Research
The Monroe Lab is driven by questions, not by any single organism. We are motivated by emerging biological discoveries now made possible by modern genomics — unexpected phenomena like dramatic variation in mutation rates, unusual chromosome biology, massive repetitive sequences, segregation distortion, and other genomic oddities that we finally have the tools to explain.
We began our work using Arabidopsis thaliana as a model system for understanding DNA repair biology and mutation. Modern sequencing now lets us scale these questions across a growing diversity of plant systems. We have active projects in:
Arabidopsis thaliana · cassava (Manihot esculenta) · pistachio (Pistacia vera) · walnut (Juglans regia) · diploid wheat (Triticum monococcum) · rice (Oryza sativa) · celery (Apium graveolens) · alfalfa (Medicago sativa) · almonds (Prunus dulcis) · black cottonwood (Populus trichocarpa)
We are entering an era where genomics can answer genuinely surprising questions about genome evolution and variation. We follow the science wherever it leads — including into some unexpected corners of genome biology.
Mutation Biology and Rate Variation
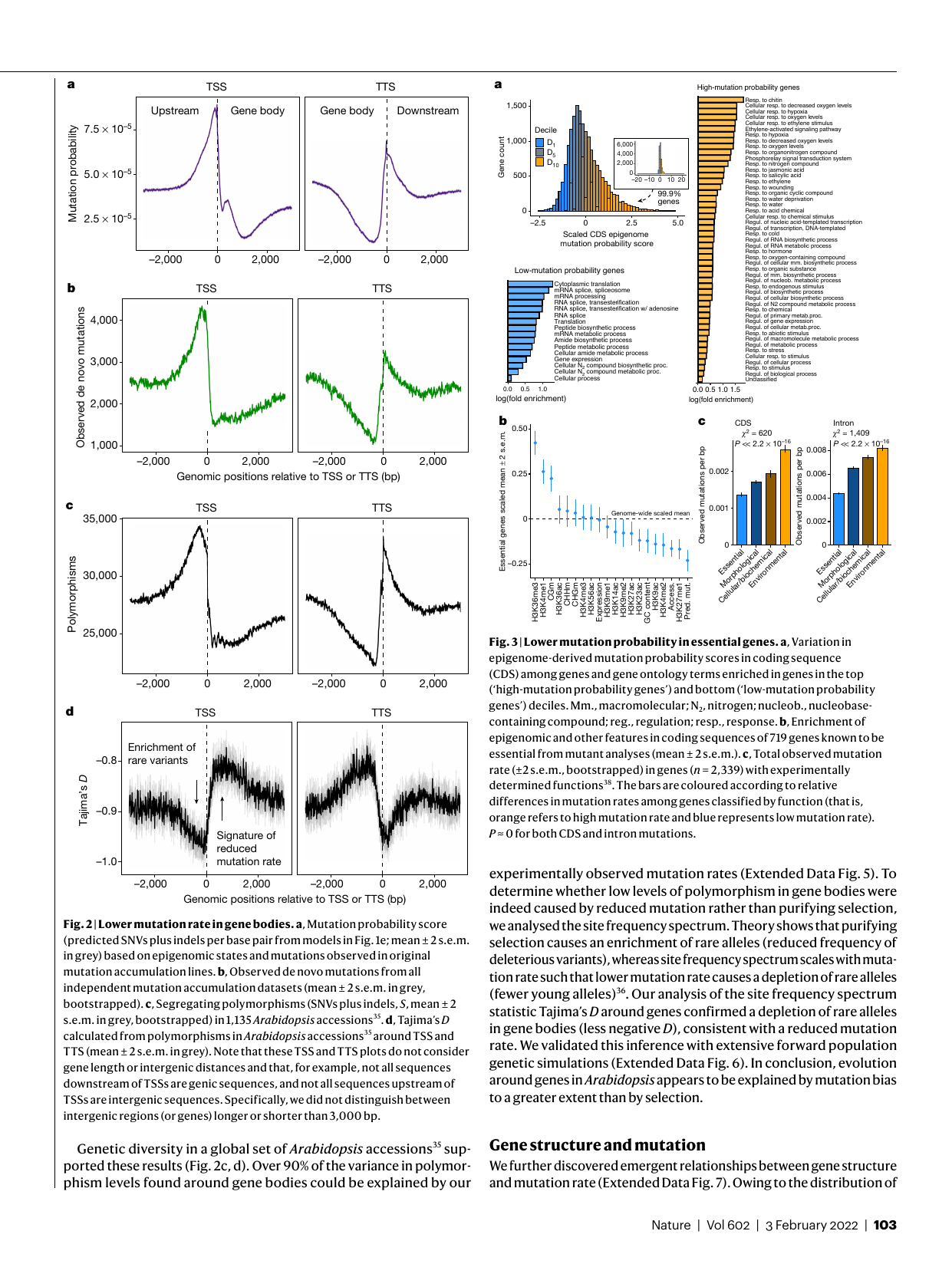
Mutations are the ultimate source of all genetic variation, yet mutation rates vary orders of magnitude across the genome, among cell types, and across species. We use ultra-accurate sequencing technologies — including Duplex Sequencing — to characterize mutation rates and spectra at unprecedented resolution. A central question is why mutation rates are systematically lower in functionally important regions of the genome, and what molecular mechanisms underlie this pattern.
Current projects include:
Epigenome-recruited DNA repair and mutation rate reduction in plants
Somatic mutation dynamics and deleterious variation in clonal walnut and pistachio cultivars
Mutation bias and natural selection in Arabidopsis thaliana
Mechanisms of mutation rate variation across the plant kingdom
Chromatin Regulation and DNA Repair
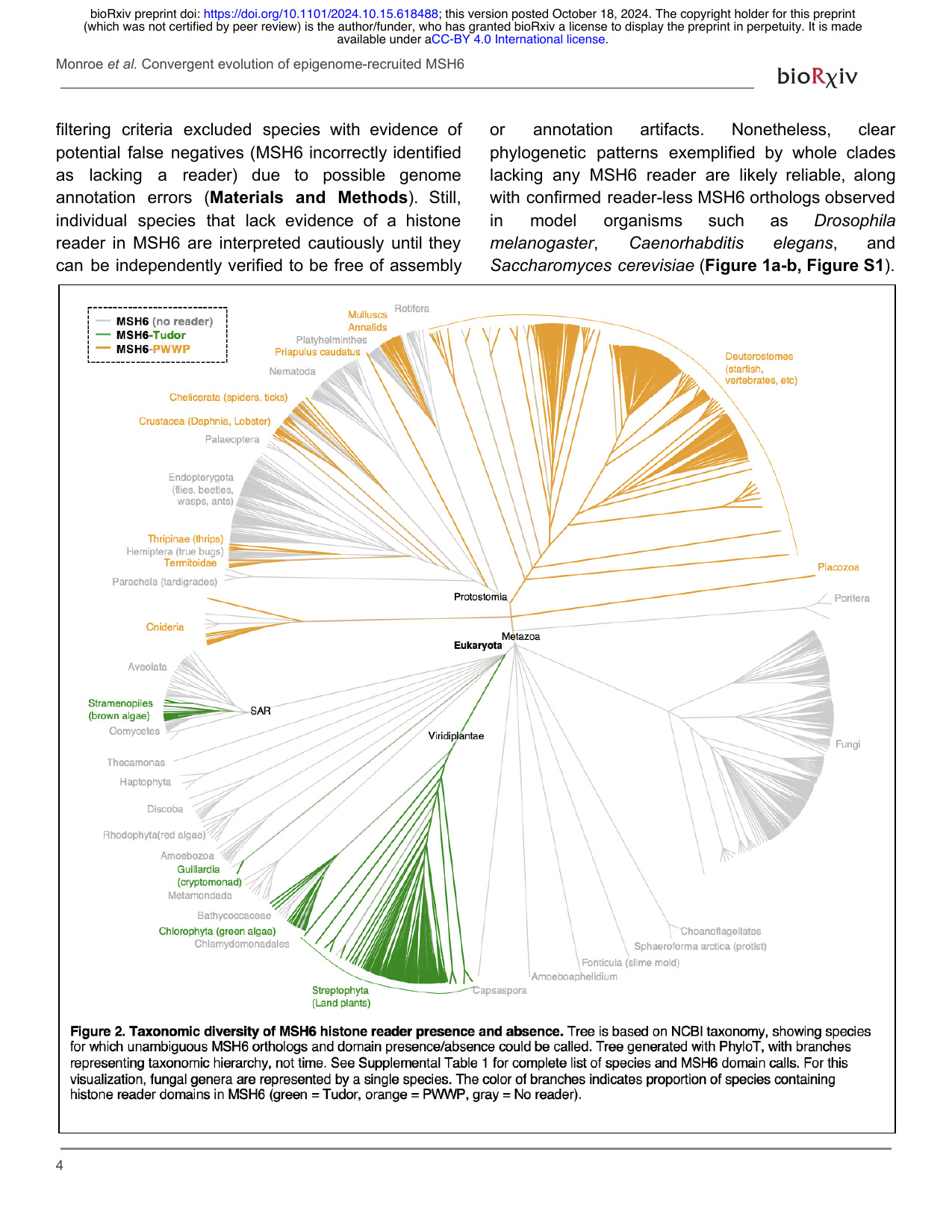
A key discovery from the lab is that the histone mark H3K4me1 directly recruits DNA repair proteins in plants, providing a mechanistic explanation for why mutation rates are lower in gene-rich, H3K4me-enriched regions of the genome. We are extending this work to understand how chromatin-targeted DNA repair has evolved independently across the tree of life and how these mechanisms shape the landscape of genetic variation within and among species.
Current projects include:
H3K4me-associated hypomutation in plants
Convergent evolution of epigenome-recruited DNA repair across the tree of life
Chromatin features predicting intron architecture and mutation rates
Genome Dynamics and Structural Variation
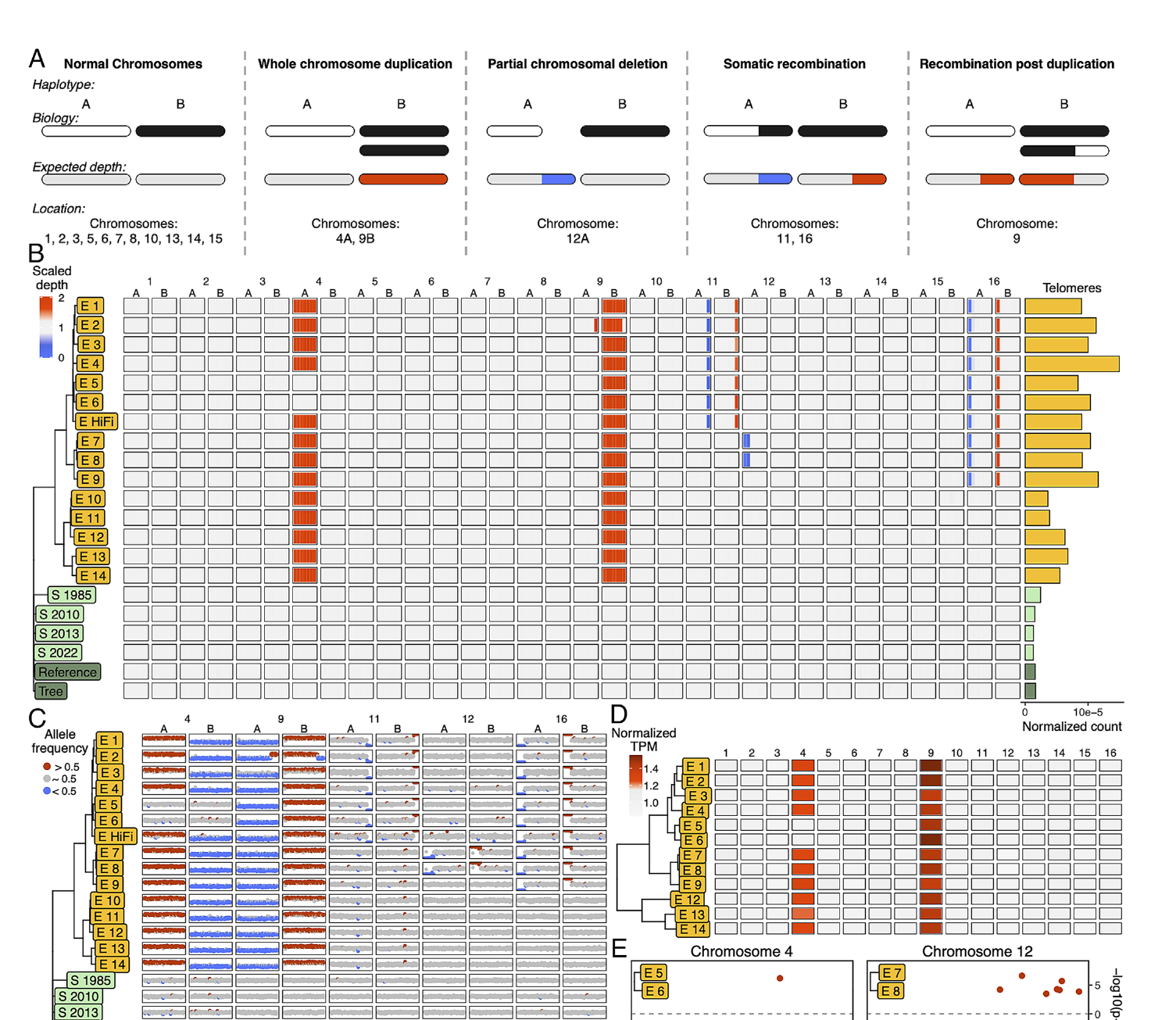
Plants harbor extraordinary structural variation — inversions, transposable elements, copy number variants, and pan-genome loci present in some individuals but absent in others. We use long-read sequencing and comparative genomics to characterize this variation and understand how it contributes to phenotypic diversity, reproductive biology, and local adaptation across plant populations and species.
Current projects include:
Pistachio pan-genome and multi-genome reference
Comparative genomics of repeat-rich and structurally complex plant genomes
Segregation distortion and unusual chromosome biology in alfalfa and other polyploids
Functional GWAS and Loss-of-Function Burden Testing
We leverage natural loss-of-function alleles to infer gene function at population scale — an approach we call functional GWAS or loss-of-function burden testing. By identifying which genes, when disrupted, produce detectable phenotypic effects across individuals, we can discover gene functions systematically and without prior functional annotation. This scales to any species with population-level genomic data and provides direct targets for breeding, gene editing, and understanding evolution.
This work builds on our foundational studies of adaptive loss-of-function in plants and extends them into crop systems where understanding which genes matter — and how — has direct practical value.
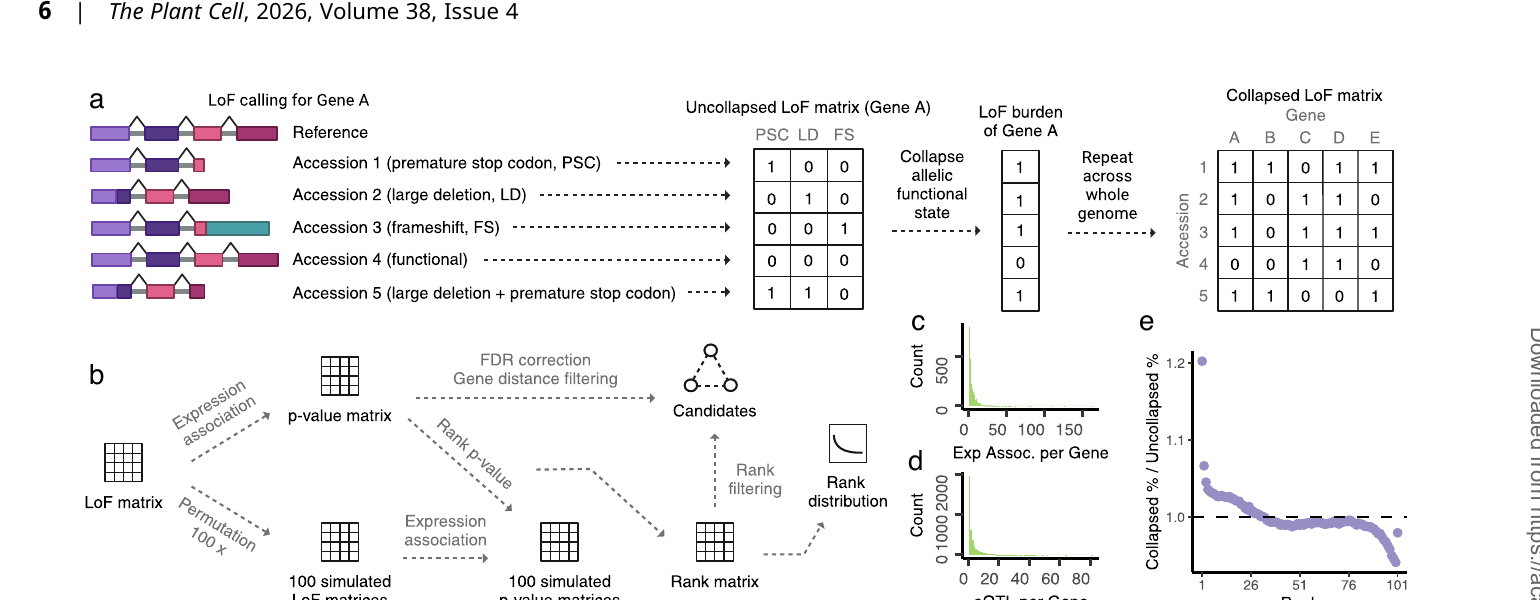
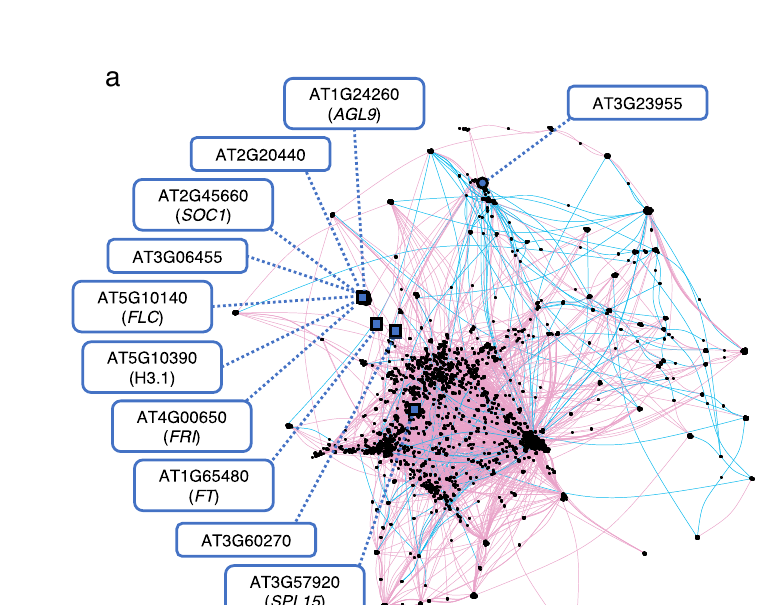
Applied to Arabidopsis thaliana, this framework recapitulates the canonical flowering-time regulatory network as an LoF-expression association graph — centered on FRIGIDA (FRI), FLC, and their interactions with SOC1, SPL15, AGL9, and histone variant H3.1 — demonstrating that LoF burden testing recovers real biology from natural variation alone.
Current projects include:
Loss-of-function burden testing in cassava (Manihot esculenta) landraces
Loss-of-function variation in diploid wheat (Triticum monococcum)
Scalable frameworks for functional gene discovery across crop species
Applied Genomics and Crop Improvement
Many of the world’s crop species remain genomically understudied. We build genomic resources — reference genomes, pan-genomes, population-scale sequencing datasets — for these species and use them to study the genetic basis of agriculturally important traits, climate adaptation, and fundamental questions about genome evolution.
Species we have worked with include pistachio (Pistacia vera), walnut (Juglans regia), cassava (Manihot esculenta), celery (Apium graveolens), diploid wheat (Triticum monococcum), rice (Oryza sativa), alfalfa (Medicago sativa), almonds (Prunus dulcis), and black cottonwood (Populus trichocarpa). Projects range from understanding mutation histories of clonal cultivars to assembling reference genomes for breeding and fundamental science.

Current projects include:
Pistachio pan-genome, reference genome, and kernel development
Cassava population genomics and climate adaptation in Colombian landraces
Climate adaptation and somatic mutation in Populus trichocarpa
Genome assembly and annotation for celery, wheat, and other crop species
Our work is supported by:
- National Science Foundation
- USDA
- California Pistachio Research Board
- Foundation for Food and Agricultural Research
- Gates Foundation
- Human Frontier Science Program
- UC Davis Department of Plant Sciences
- UC Davis Venture Catalyst